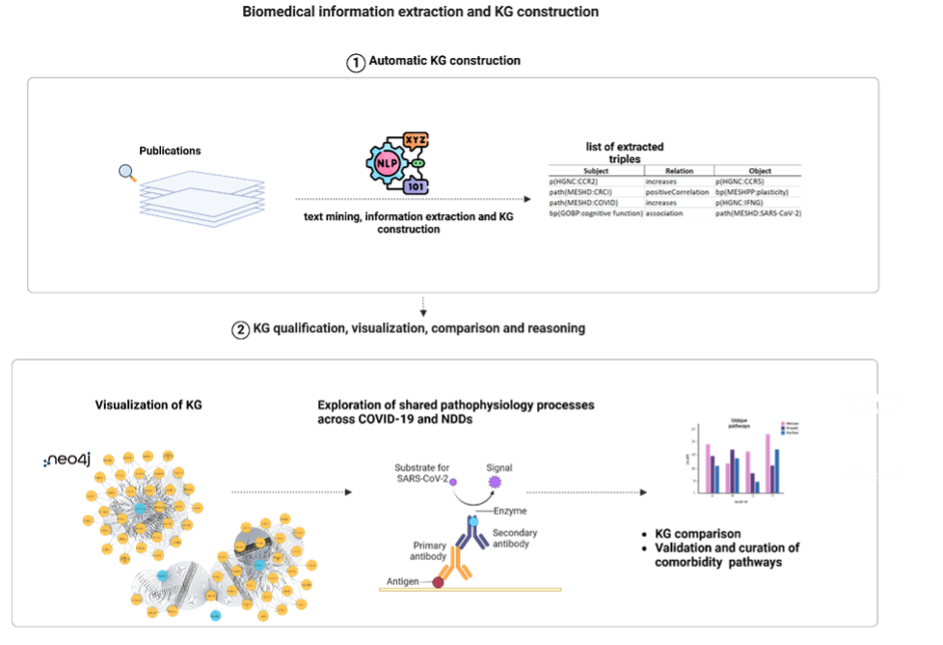
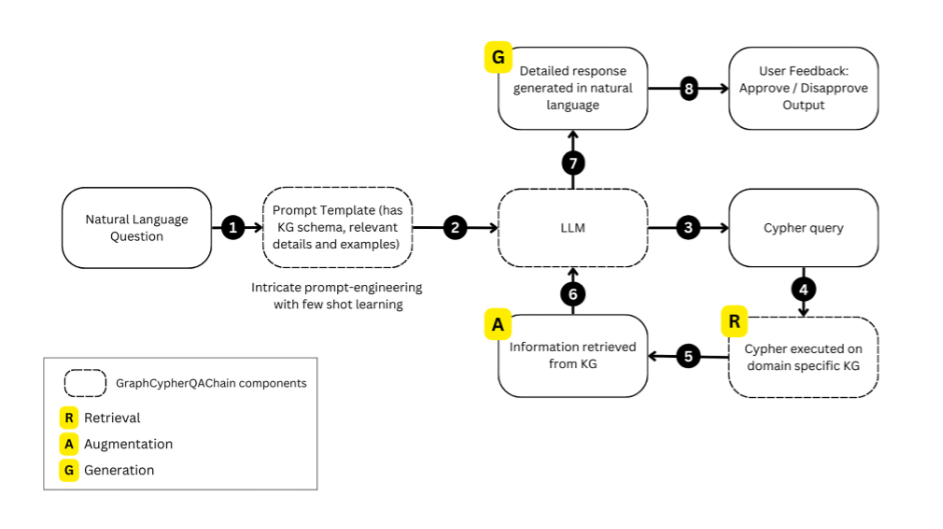
Overview
Lead: Fraunhofer SCAI
Partners: University of Luxembourg, Kairntech
WP2 builds and operates the data and knowledge backbone of COMMUTE, integrating heterogeneous data and structured biomedical knowledge to study shared mechanisms between COVID‑19, Alzheimer’s disease (AD) and Parkinson’s disease (PD).
Key activities include:
- Serving as the central hub for data harmonization, curation and annotation across all partners.
- Integrating EHRs, organoid / wet‑lab data, environmental data, and literature‑derived knowledge graphs into a coherent ecosystem.
- Establishing shared semantic frameworks so datasets and tools from different work packages can interoperate.
- Developing, updating and deploying disease knowledge graphs (e.g. Parkinson’s Disease map, NeuroMMSig) in a Neo4j graph database.
- Using AI/NLP‑based literature analysis and in‑silico simulations to derive and refine mechanistic hypotheses.
- Operating the COMMUTE Evidence Base, which shares testable hypotheses, candidate biomarkers, ML/AI models and selected shareable datasets.
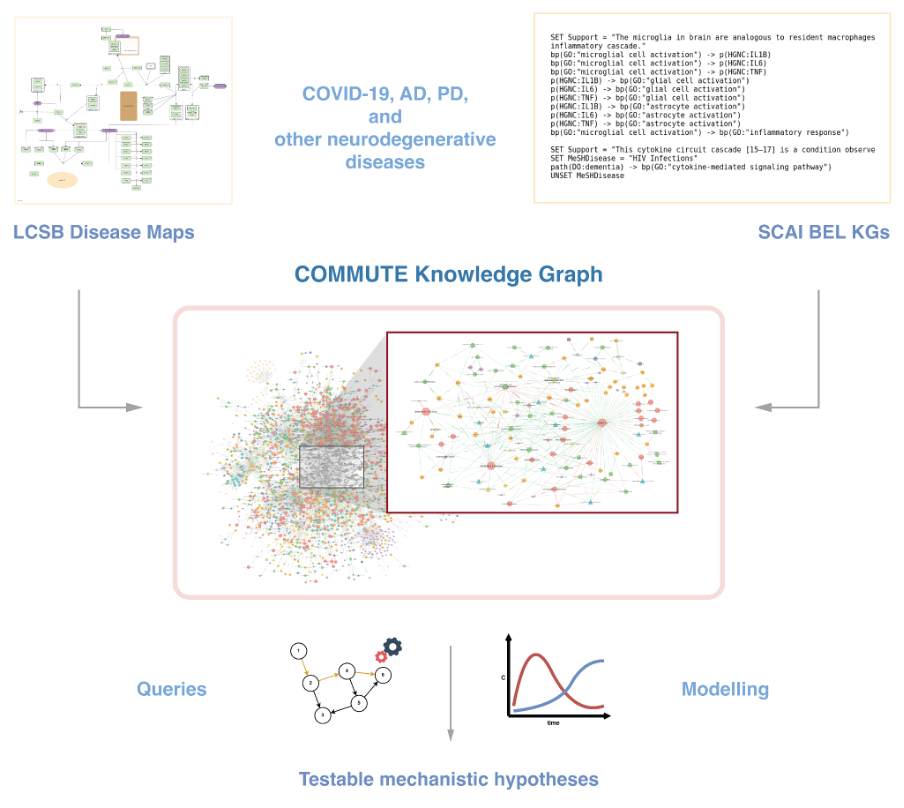
 Comorbidity Mechanisms Utilized in Healthcare
Comorbidity Mechanisms Utilized in Healthcare